DECODING THE BOOK OF LIFE: A GUIDE TO WHOLE GENOME SEQUENCING
- Related toScience & Tech
- Published on14 June 2025
 Have you ever wondered why you have your mother’s eyes or your father’s smile? The answer lies in a microscopic instruction manual hidden inside almost every cell in your body. This manual, written in a four-letter alphabet, dictates everything from your appearance to your susceptibility to certain diseases. For centuries, this “Book of Life” remained sealed. We knew it existed, but we could only guess at its contents. Today, that has changed. Thanks to a revolutionary technology called Whole Genome Sequencing (WGS), we can now read this entire book, from cover to cover, letter by letter. This is not just a scientific breakthrough; it’s a paradigm shift that is beginning to unlock the deepest secrets of our health, our history, and what it means to be human.
Have you ever wondered why you have your mother’s eyes or your father’s smile? The answer lies in a microscopic instruction manual hidden inside almost every cell in your body. This manual, written in a four-letter alphabet, dictates everything from your appearance to your susceptibility to certain diseases. For centuries, this “Book of Life” remained sealed. We knew it existed, but we could only guess at its contents. Today, that has changed. Thanks to a revolutionary technology called Whole Genome Sequencing (WGS), we can now read this entire book, from cover to cover, letter by letter. This is not just a scientific breakthrough; it’s a paradigm shift that is beginning to unlock the deepest secrets of our health, our history, and what it means to be human.
The Blueprint of Life: What Exactly is a Genome?
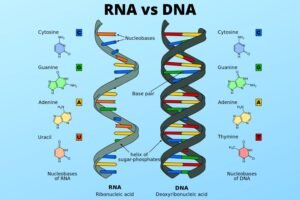 Before we dive into how we read the book, let’s understand what it is. An organism’s genome is its complete set of genetic instructions. Think of it as an incredibly detailed instruction manual. This manual is written in the language of DNA (Deoxyribonucleic acid), using just four chemical “letters”: A (Adenine), T (Thymine), C (Cytosine), and G (Guanine).
Before we dive into how we read the book, let’s understand what it is. An organism’s genome is its complete set of genetic instructions. Think of it as an incredibly detailed instruction manual. This manual is written in the language of DNA (Deoxyribonucleic acid), using just four chemical “letters”: A (Adenine), T (Thymine), C (Cytosine), and G (Guanine).
In humans, this manual contains about 3.2 billion letters, neatly packaged into structures called chromosomes. You can think of chromosomes as the chapters of the book. Within these chapters are specific sections called genes, which are like individual recipes or instructions. One gene might hold the recipe for producing insulin, while another might have the instructions for building the proteins that determine your blood type. The entirety of these chapters and recipes constitutes your genome.
Reading the Entire Book: Understanding Whole Genome Sequencing
For a long time, our ability to read this book was limited. We could perhaps read a few select recipes (genes) or check for specific spelling mistakes (mutations). This is what older methods like DNA profiling or SNP genotyping do. Whole Exome Sequencing, a more advanced technique, allows us to read all the recipe sections (the exome), which make up about 1-2% of the entire book.
Whole Genome Sequencing (WGS), however, is in a league of its own. As the name suggests, it is the process of determining the precise order of all 3.2 billion letters in an organism’s genome in a single go. It reads not just the recipes (genes) but also the introductions, the indexes, the spaces between the paragraphs, and all the seemingly “junk” DNA that we are only now beginning to understand is critically important. It gives us the complete, unabridged story.
 A Journey Through Time: The Quest to Sequence the Genome
A Journey Through Time: The Quest to Sequence the Genome
The ability to read the book of life didn’t happen overnight. It’s the culmination of decades of scientific endeavour.
The journey began in the 1970s with slow, manual sequencing methods. The first complete genome ever sequenced, in 1976, belonged to a tiny virus. The scale of the challenge grew with each milestone.
In 1995, scientists achieved a landmark by sequencing the entire genome of a free-living organism, the bacterium Haemophilus influenzae. This was followed by the first complex organism (a yeast) in 1996, and the first multicellular animal (a nematode worm) in 1998.
The true watershed moment came with the Human Genome Project (HGP) in the field of genome sequencing. Launched in 1990, this monumental international collaboration set out to do what seemed impossible: to sequence the entire human genome. After 13 years of relentless work, a draft of the human genome was published in 2003, with a more complete version following in 2004. The HGP was a scientific moonshot, and its success opened the floodgates for the genomic revolution we are witnessing today.
The ‘How-To’: A Glimpse into the Sequencing Process
So, how do scientists read 3.2 billion letters of DNA? The most common modern approach is a form of “shotgun sequencing” combined with Next-Generation Sequencing (NGS) technology.
Imagine taking the entire instruction manual and putting it through a shredder, creating millions of tiny, overlapping snippets of text. Now, imagine a machine that can read all these millions of snippets simultaneously. That’s essentially what NGS platforms do. They rapidly read these small fragments of DNA. The final, and perhaps most challenging, step is to use powerful computers to look at the overlapping ends of these snippets and piece them back together in the correct order, reconstructing the entire book.
Newer technologies, like long-read sequencing, are also emerging. While perhaps less accurate letter-by-letter, they read much longer snippets of text, making it easier to solve the puzzle, especially in parts of the book with lots of repetitive sentences (complex regions of the genome).
Unlocking the Secrets: What Can We Do with a Whole Genome?
Having the complete instruction manual for an individual opens up a world of possibilities, particularly in medicine and healthcare.
A New Era of Personalised Medicine: For decades, medicine has largely followed a one-size-fits-all approach. WGS is changing that. By understanding an individual’s unique genetic makeup, doctors can predict how they might respond to certain drugs, allowing them to choose the most effective treatment and avoid adverse reactions.
Diagnosing the Undiagnosable: For families affected by rare genetic diseases, the diagnostic odyssey can be long and heartbreaking. WGS can provide answers where other tests fail. By scanning the entire genome, it can pinpoint the single “typo” among billions of letters that is responsible for a mysterious condition, providing a diagnosis and paving the way for potential treatments.
Transforming Cancer Care: Cancer is a disease of the genome. Tumors develop because of mutations in a cell’s DNA. By sequencing the genome of a tumor, doctors can understand exactly what went wrong and choose targeted therapies designed to attack cancer cells with those specific mutations. Furthermore, a technique called “deep whole genome sequencing” can analyze tiny fragments of tumor DNA circulating in the bloodstream (ctDNA), offering a non-invasive way to detect cancer early, monitor treatment effectiveness, and understand how the cancer is evolving.
Public Health and Epidemics: In the face of an epidemic, WGS is a critical tool for public health. By sequencing the genome of a virus or bacterium, scientists can track how it is spreading, how it is mutating, and develop vaccines and treatments to combat it. The rapid sequencing of the SARS-CoV-2 virus was instrumental in the global response to the COVID-19 pandemic.
The Indian Chapter: The Genome India Project (GIP)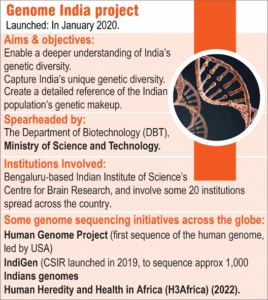
Launched in 2020 by the Department of Biotechnology (DBT), the Genome India Project (GIP) is a nationwide scientific initiative aimed at decoding the genetic makeup of India’s diverse population. In collaboration with 20 research institutions across the country, this genome sequencing project aims to build a comprehensive catalogue of genetic variations unique to different Indian communities. This is crucial for understanding how genes influence health, disease, and traits among Indians.
So far, the project has collected around 20,000 DNA samples from 83 different population groups, representing India’s broad genetic diversity. Out of these, the genomes of 10,000 individuals have already been successfully sequenced during the first phase. This data will help create a reference genome specifically tailored for the Indian population – a vital tool for improving healthcare research and diagnosis. All this data is stored at the Indian Biological Data Centre (IBDC) in Faridabad, which is India’s first national data repository for life science research. IBDC has also launched a new protocol called the Framework for Exchange of Data (FeED), ensuring transparent, fair, and secure sharing of genetic information under national biotech guidelines.
Genome sequencing has many far-reaching applications. In the field of healthcare, it allows scientists to study genetic disorders, track the origin of diseases, and develop precision medicine suited to an individual’s DNA. In agriculture, it helps in improving crop varieties and livestock health. It also supports biodiversity conservation by helping scientists document and study various species.
10,000 Indian genomes sequenced in Phase I; stored at IBDC, Faridabad.
83 diverse population groups covered; 20 institutions collaborated.
FeED Protocol launched to enable ethical, secure data exchange.
However, there are some challenges as well. There are concerns about data privacy, ethical usage, and a lack of strong regulations, especially as many Indian genetic samples are sent abroad for testing. Additionally, the high cost of genome sequencing and fragmented data systems limit its accessibility and use in public health planning. But with the rise of newer technologies like Next-Generation Sequencing (NGS), these challenges are gradually being addressed. NGS makes genome decoding faster, cheaper, and more efficient, bringing us closer to a future where personalised medicine and preventive healthcare become a reality for all.
The Ethical Maze: Navigating the Challenges of Genomic Data
The power of WGS also brings with it significant ethical and social challenges that we must navigate carefully.
- Data Privacy and Security: Your genome is your most personal information. How do we ensure this data is stored securely and protected from misuse? The risk of data breaches is a serious concern.
- Genetic Discrimination: Could insurance companies or employers use your genetic information to discriminate against you? Many countries are grappling with this question, with some, like the US, having enacted laws like the Genetic Information Nondiscrimination Act (GINA) to prevent this.
- Informed Consent: The results of a genome sequencing can reveal life-altering information, such as the risk of incurable diseases or even details about one’s paternity. Ensuring that individuals give truly informed consent and receive proper counselling to understand the psychological and social implications is crucial.
- Equity and Accessibility: While the cost of sequencing has plummeted from billions of dollars for the Human Genome Project to a few hundred dollars today, it is still not universally accessible. There is a risk that genomic medicine could widen the gap in healthcare access between the rich and the poor.
Conclusion: The Future is Written in Our DNA
Whole Genome Sequencing has transformed biology from a science of observation to a science of information. It has taken us from guessing at the contents of the Book of Life to having the entire text at our fingertips. The journey from a monumental research effort to a practical clinical tool has been nothing short of breathtaking. As the technology continues to evolve and costs continue to fall, WGS promises a future where medicine is more predictive, personalized, and effective than ever before. However, as we unlock the secrets of our own code, we must also embrace the responsibility of being wise and ethical stewards of this profound knowledge. The future of health is written in our DNA, and we are, for the first time, learning how to read it.
Basics of Genome & WGS
- Genome = Complete set of genetic instructions (DNA) in an organism.
- Written using 4 letters: A, T, G, C.
- Human genome = ~3.2 billion letters, organized in chromosomes and genes.
Whole Genome Sequencing (WGS)
- WGS reads the entire genome—not just genes but all regions, including “junk DNA”.
- More comprehensive than DNA profiling or Whole Exome Sequencing.
- Uses Next-Generation Sequencing (NGS) for faster and cheaper decoding.
How WGS Works
- The genome is cut into small fragments.
- NGS machines read these simultaneously.
- Powerful computers reassemble the full genome.
Historical Milestones
- 1976: First genome sequenced (virus).
- 1995–1998: First free-living organism, yeast, and nematode sequenced.
- 1990–2003: Human Genome Project (HGP) successfully sequenced human genome.
Applications of WGS
Personalised Medicine – Predicts drug responses and avoids side effects.
Rare Disease Diagnosis – Finds exact genetic causes of undiagnosed conditions.
Cancer Treatment – Identifies mutations for targeted therapies and early detection through ctDNA.
Public Health – Tracks virus mutations (e.g., SARS-CoV-2), aids in vaccine development.
Genome India Project (GIP)
- Launched by DBT in 2020.
- Aim: Decode genetic diversity across Indian population groups.
- 10,000 genomes sequenced, data stored at IBDC, Faridabad.
- Uses FeED protocol for secure and ethical data exchange.
- Involves 20 institutions and 83 population groups.
Additional Benefits
Enhances research in healthcare, agriculture, and biodiversity conservation.
Ethical Challenges
- Data Privacy – Genome data is highly sensitive and must be protected
- Genetic Discrimination – Possible misuse by insurers or employers.
- Informed Consent – People must understand the implications of sequencing results.
- Accessibility Gap – High cost and digital divide may limit access to genomic healthcare
Loved this article? Go to Learning EDGE+ Page↗️

